Developer Guide
The preceding documentation provides a more conceptual understanding of the VA-Spec for a broad audience - covering its content, modeling principles, and general utility. This section provides more technical guidance to support modelers and data engineers who will be authoring VA Profiles, or implementing them in data exchange systems.
Profile Definition Mechanisms
We have previously described two categories of profiles in the VA-Spec, which are authored using different mechanisms:
Base Profiles: use a subclassing mechanism to define Proposition and Study Result profiles as VA base classes.
Community Profiles: use a schema-composition mechanism to define Statement and Evidence Line profiles as constraints on top of core class definitions.
For Proposition and Study Result Base Profiles, a subclassing mechanism is required to rename and add additional attributes - including qualifier and data item fields used to collect domain-specific information in these profiles
For Statements and Evidence Lines, any domain-specificity is specified in the Propositions these objects encapsulate, so there is no need to define formal subclasses here. However, VA-Spec includes Community Profiles of these classes that constrain certain attribute values to align with the conventions of a particular community guideline - and here schema composition is sufficient to define these restrictions. This approach to profile definition reduces the number of classes that need to be created, managed, and parsed when creating and validating VA data.
The diagrams below illustrate where subclass- and composition-based mechanisms are applied to define each profile included in the VA-Spec. Top to bottom, the increasingly dark colors reflect the increasing domain-specificity of the models.
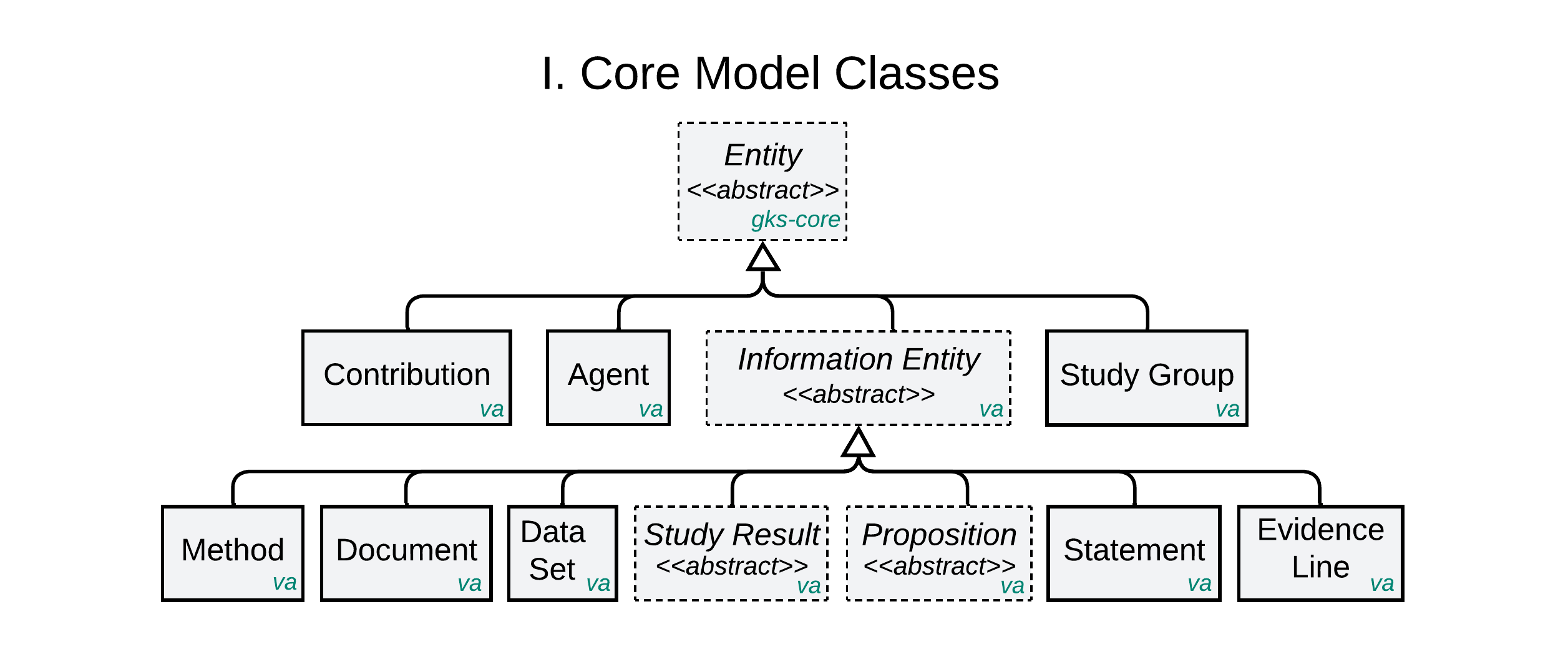
The Core Data Model consists of the domain-agnostic classes above. Concrete classes can be used to capture data directly. Abstract classes must first be ‘specialized’ through subclassing. Note that some classes in the model are imported from gks-core, vrs, and cat-vrs models, as indicated by annotations in green which indicate the GKS specification in which each is defined.
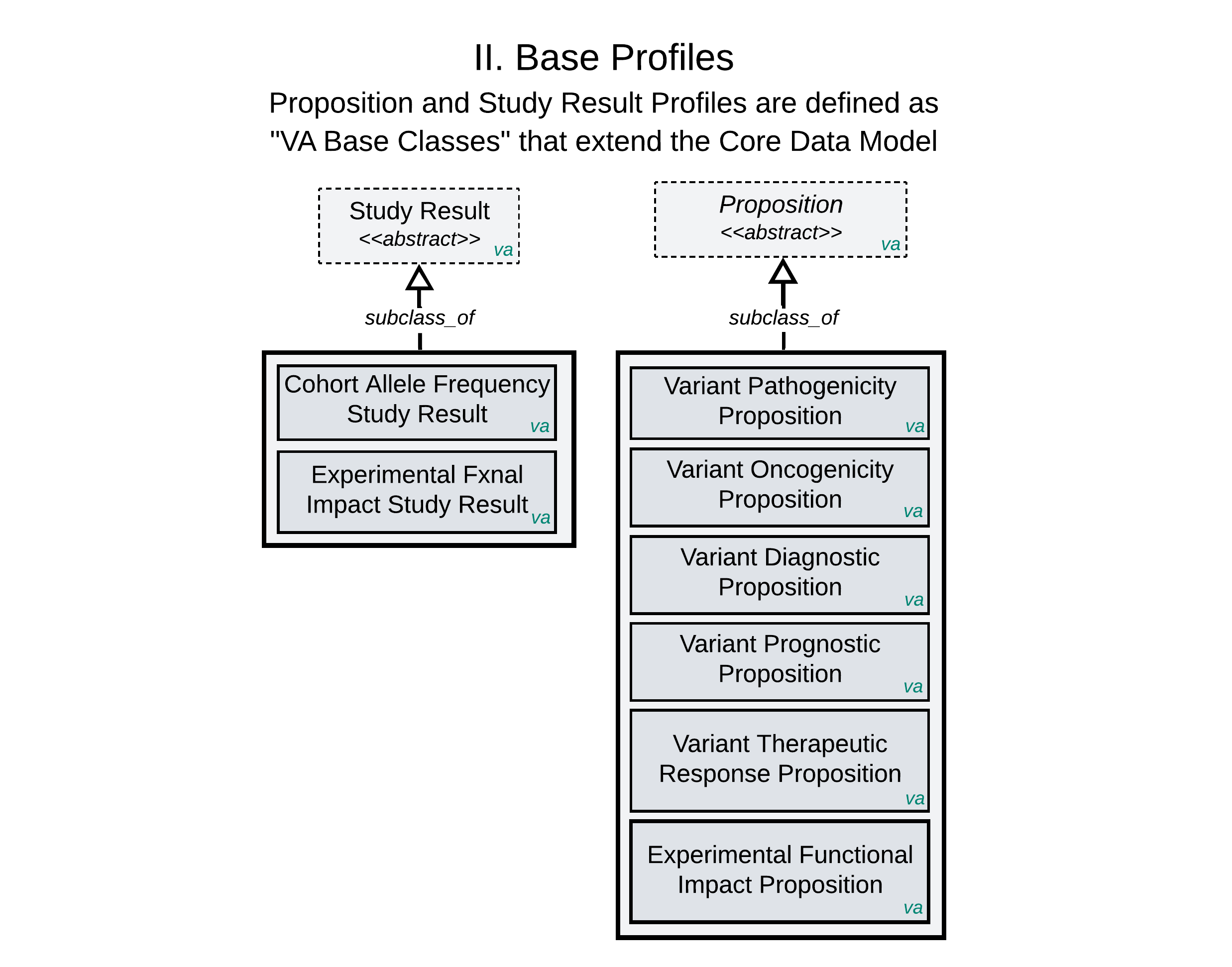
The Proposition and Study Result Base Profiles above are defined using a subclassing mechanism, creating formal “VA Base Classes” that extend the Core Data Model. The specific syntax for this authoring mechanism leverages features outside the native JSON Schema language, as illustrated in the Proposition profile example here.
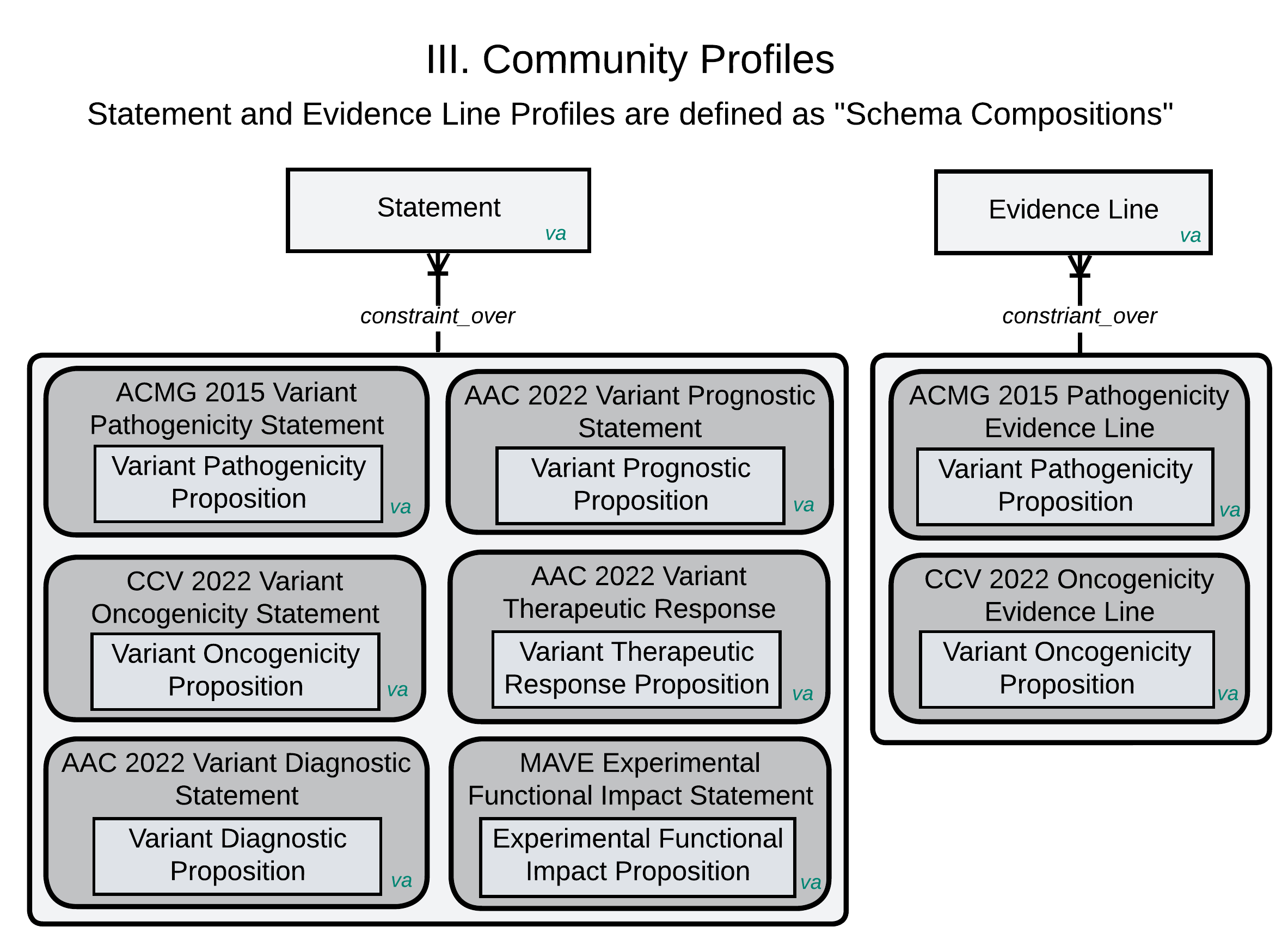
The Statement and Evidence Line profiles above are defined as “Schema Compositions” using a constraint-based mechanism. These profiles represent sub-schema, rather than sub-classes in the VA Model. The domain-specificity of these profiles is defined in the Proposition profiles they encapsulate, as diagrammed. Constraints may be added to restrict certain attributes to align with terminological conventions of a particular community guideline (e.g. ACMG-2015, AAC-2017, CCV-2022). The specific syntax for this authoring mechanism is illustrated in the Statement profile example here.
Finally, this diagrammed data example provides a nice visualization of how Core Model classes and profiles defined using these different mechanisms are used together to represent real data. Styling conventions in the diagram indicate the type of model that specifies each object in the example (Core Class, Base Profile, Community Profile).
Custom Profile Development
Representation of a particular type of Statement or Evidence Line using the VA-Spec does not always require a VA Profile to be specifically defined for it.
Custom Profiles are Statement or Evidence Line models that are defined de novo, to support a specific implementation use case where data cannot be made to conform to a particular guideline-based Community Profile.
This section describes why these are useful, and how to create them.
Why they are useful:
The Statement and Evidence Line Community Profiles included in version 1.0 of the VA-Spec are there to support data providers pursuing strict alignment with a particular community guidelines.
Implementers who do not seek such alignment can build their own schema for Statements or Evidence Lines to report on any of the knowledge types specified in VA Base Proposition profiles.
For example, a project that aims to represent some of the messier data in ClinVar where values for key fields bound to ACMG-specific enumerations in the existing Variant Pathogenicity Statement profile - and doesn’t want to use Extensions to capture this data - can define a custom Pathogenicity Statement Profile from core Statement and Evidence Line classes that applies constraints specific to its data.
The process is straightforward - e.g. to create a custom Statement profile for pathogenicity classification data not based strictly aligned with ACMG terminology:
Start with the core Statement class
Bind its
propositionattribute to the VariantPathogenicityProposition base profile classUse other core Statement attributes and related core classes to represent additional information about the Statement (e.g. strength, classification, methods, etc) - defining additional constraints or enumerations as desired using the Composition-Based Profiling Mechanism described above.
Use the base Statement reference implementation to create and validate compliant data.
This simple data example illustrates application of this approach to create a custom, non-ACMG-compliant representation of a pathogenicity statement.