Profiles
Important
This overview is meant to provide a conceptual understanding of how profiling works. The technical details behind how profiles are specified and used are described in the Developer Guide section.
Profiling Tasks
As noted, the VA-Spec provides Statement, Study Result, Evidence Line, and Proposition Profiles that specialize these core classes to support a specific domain of variant knowledge (e.g. pathogenicity), and/or support conventions of a particular community guideline (e.g. ACMG-2015).
The table below describes the different profiling tasks supported in v1.0 of the VA-Spec, with examples based on definition of ACMG-aligned Variant Pathogenicity profiles. These are illustrated graphically in the figure that follows.
Profiling Task |
Example |
|---|---|
Define a domain- or community-specific version of a Core Model class |
Specialization of the |
Import and reference classes for domain entities that variant knowledge is about |
The |
Constrain attributes to take a more specific type of value (possible renaming them in the process) |
The |
Define value sets and binding them to select attributes. |
The |
Define a new attribute to capture domain-specific information in a profiled class |
The |
Refine cardinality of select attributes |
The |
Profiling Example
The diagram below illustrates at a conceptual level some of the profiling steps applied to the core Statement and Proposition classes, to create models supporting ACMG-based Variant Pathogenicity Statements.
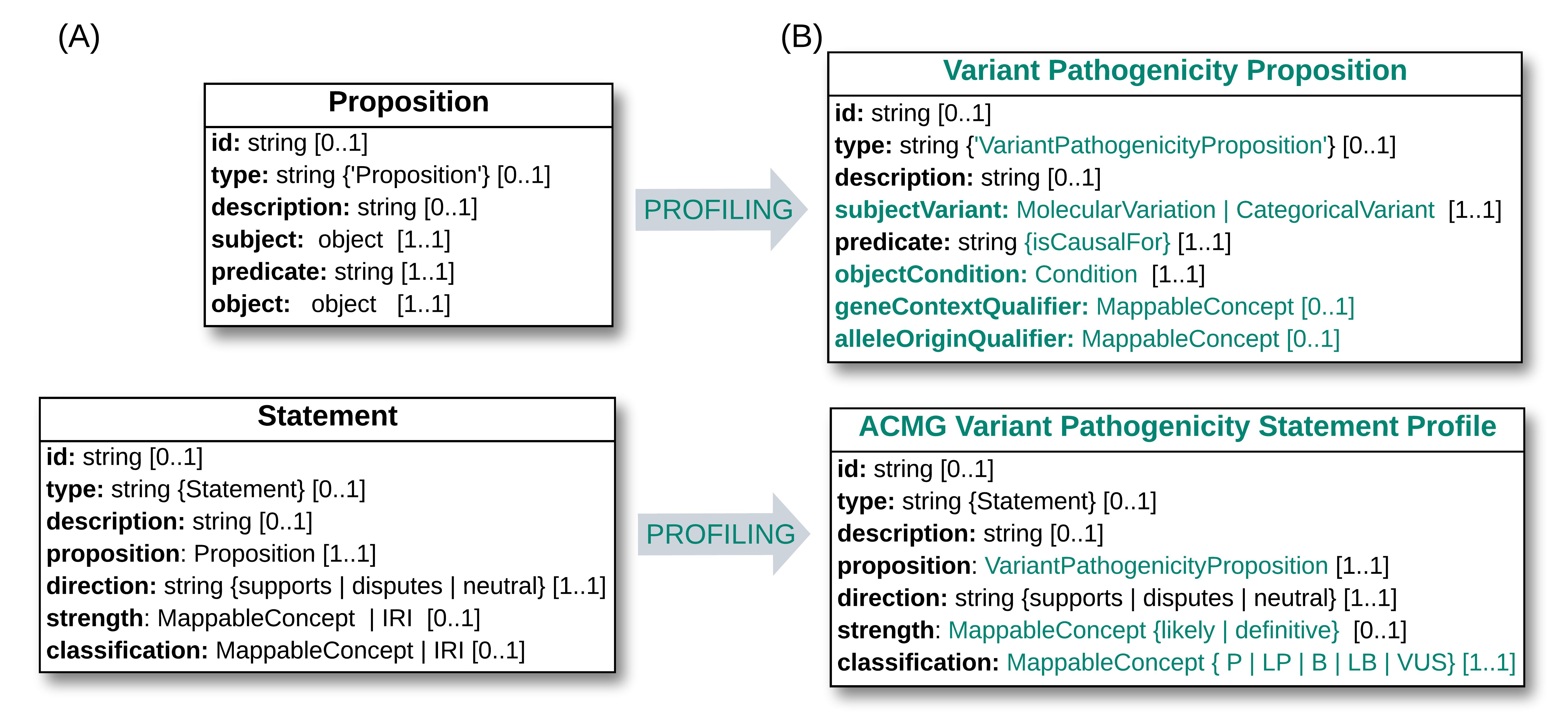
ACMG-based Variant Pathogenicity Profiles.
(A) Core Proposition and Statement classes, showing a subset of their attributes. (B) ACMG-based Variant Pathogenicity profiles derived from these core classes, with profiling specializations in green. Text in curly braces are enumerations, which in some cases are nested inside fields of a MappableConcept. The actual VA-Spec v1.0 schema for these profiles are here and here.
This data example illustrates application of these two profiles to represent a simple Variant Pathogenicity Statement.
Future versions of the VA-Spec will include a more formal specification and tooling support for executing these tasks and validating they were performed correctly.