ACMG Variant Pathogenicity Statement Example (with Evidence)
Description:
The data below builds on the simple ClinVar-GKS example described here, embellishing its base ClinVar record with additional evidence to demonstrate richer structures the Variant Pathogenicity Statement (ACMG 2015) profile can support.
Specifically, it stitches together several simpler Statement, Study Result, and Evidence Line data examples from the test fixtures directory, to reveal how these objects can be combined to build the rich evidence and provenance structure below.
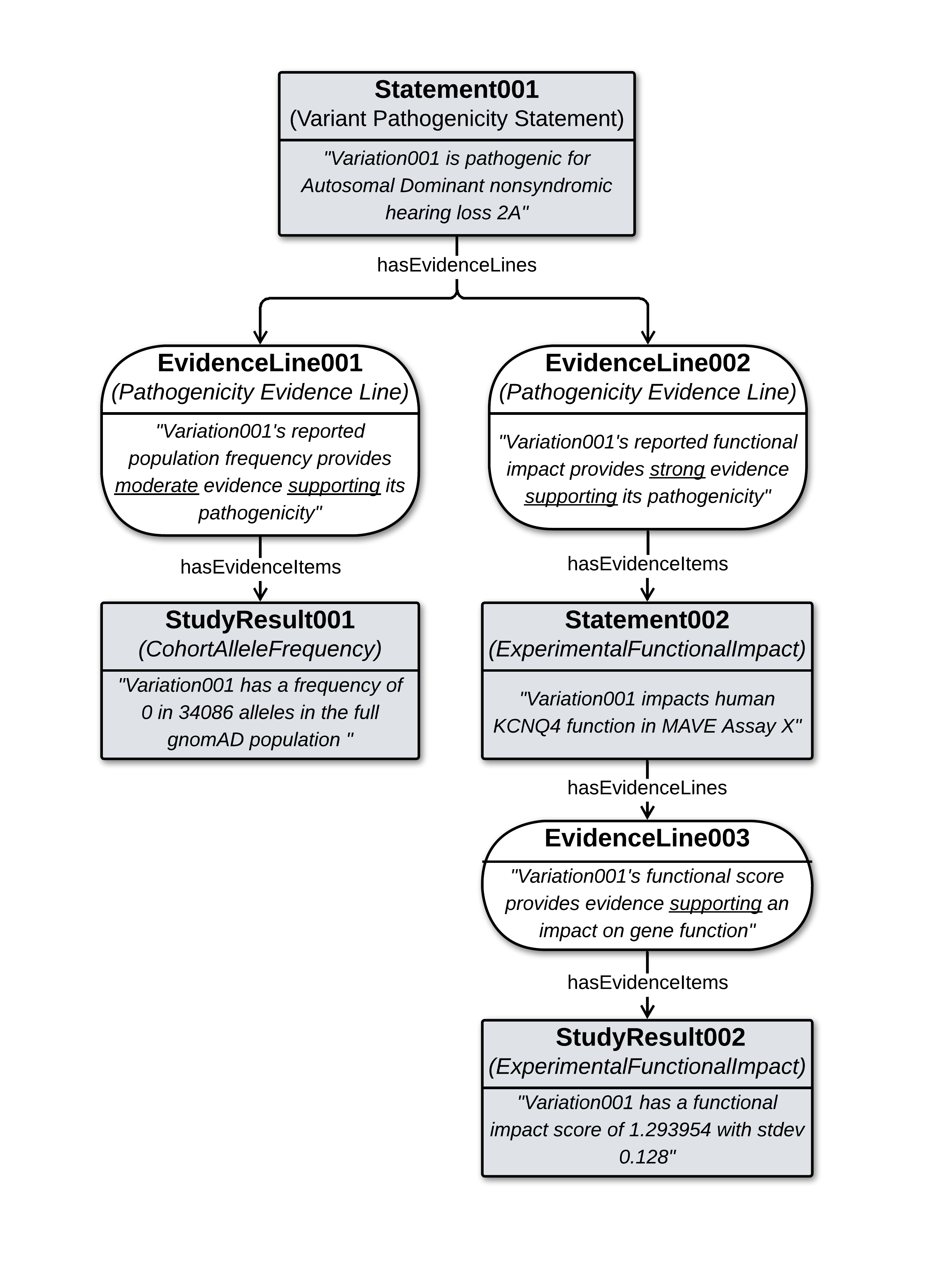
High Level Structure of the Data Example
Legend: A root Pathogenicity Statement is supported by Evidence Lines based on a Cohort Allele Frequency Study Result from gnomAD, and a Functional Impact Statement from MAVE DB, which itself is supported by a Functional Impact Study Result. Boxes represent objects comprising the central axis of the data, with italicized text indicating what each object reports to be true.
Such structures can represent the full details of how evidence is interpreted to build up support for higher order assertions of variant knowledge - e.g. here how functional data from a study result supports a study-specific conclusion about the functional impact of a variant, which is interpreted as ‘strong’ evidence ‘supporting’ for the variant’s possible pathogenicity, and assessed as one argument supporting an ACMG-based pathogenicity classification of the variant.
A few additional notes about this example:
Comments in the yaml are provided to help readers better understand the structure, semantics, and utility of the data in the example.
Some identifiers not present in the source test fixture data were created for purposes of identifying and cross-referencing objects in this aggregate example (these are all prefixed with the string ‘ex:’).
Note that the variant subject of each Statement and Study Result objects is reported as the same, generic variation for simplicity (ex:Variant001). In reality these objects may describe subtly different variants that all map to each other in some way (e.g. a protein-level variant in the Functional Impact objects, a genomic-level variant in the Allele Frequency objects, and a Categorical Variant that covers both of these contextual variants in the Pathogenicity Statement and its direct Evidence Lines). Nuances around how variant subjects of Statements and those described by supporting evidence is a separate and complex topic addressed here.
The example omits full representations of these VRS and CatVRS Variation objects - as these are large structures that are the remit of other GKS Specifications.
Data:
Note
We recommend opening this example side-by-side with the figure above, and tracking how the data reflects the diagrammed structure and semantics.
ex.Statement001:Based on the ClinVar record SCV000778434.1
id: ex:Statement001
type: StatementFormal type in the model is 'Statement', but the data aligns with the "Variant Pathogenicity Statement (ACMG 2015)" Community Profile.
proposition:a Proposition object captures the possible fact assessed by the Statement, using a subject, predicate, object, qualifier (SPOQ) semantic modeling pattern.
id: ex:Proposition001the proposition here is that "NM_004700.4:c.803CCT[1] is causal for AD nonsyndromic hearing loss 2A"
type: VariantPathogenicityProposition
subjectVariant: ex:Variant001'subjectVariant' specializes the VA Core 'subject' attribute. The full representation of the NM_004700.4:c.803CCT[1] KCNQ4 variant is not included.
predicate: isCausalForthe predicate for this Statement profile is fixed at 'isCausalFor'
objectCondition:'objectCondition' specializes the VA Core 'object' attribute.
id: clinvar.trait/939this is a MappableConcept object that represents the Condition, using names/codes from existing code systems
conceptType: Disease
name: Autosomal dominant nonsyndromic hearing loss 2Athe name for the concept as assigned by the data provider
primaryCoding:holds a Coding object, where the concept is defined in the 'code' or 'name' field
code: C2677637the code from the MedGen terminology for AD nonsyndromic hearing loss 2A
system: https://www.ncbi.nlm.nih.gov/medgen/
iris:
- http://identifiers.org/medgen/C2677637
penetranceQualifier:holds a MappableConcept that reports qualifying penetrance information about the object condition (here, that the statement holds for high penetrance AD hearing loss)
primaryCoding:
code: high
system: ga4gh-gks-term:pathogenicity-penetrance-qualifiercode system here is a locally defined placeholder, until we formalize terminological standards for use in the VA-Spec
name: high
direction: supportsan enumerated string that indicates the Statement 'supports' the Proposition as true
strength:holds a MappableConcept reporting that confidence/evidence for this stated support
primaryCoding:
code: definitivethe code here is a term based on language used in the ACMG guidelines, as ACMG does not provide a formal code system for this
system: ACMG Guidelines, 2015
classification:holds a MappableConcept reporting the final ACMG classification of the subject variant to be 'pathogenic'
primaryCoding:
code: pathogenicthe code here is a term based on language in the ACMG guidelines, as ACMG does not provide a formal code system for this
system: ACMG Guidelines, 2015
contributions:a list of Contribution objects, each describing how an agent contributed to the Statement
- type: Contribution
contributor:reports who made this contribution
id: clinvar.submitter/500139
type: Agent
name: ClinVar Staff, National Center for Biotechnology Information (NCBI)
activityType:reports the type of contribution that was made (here an evaluation activity)
name: evaluated
mappings:
- coding:
code: cg000011
system: https://dataexchange.clinicalgenome.org/codes/
relation: exactMatch
date: '2015-08-20'reports when this contribution was performed
- type: Contribution
contributor:
id: clinvar.submitter/500139
type: Agent
name: ClinVar Staff, National Center for Biotechnology Information (NCBI)
activityType:
name: submitted
mappings:
- coding:
code: cg000010
system: https://dataexchange.clinicalgenome.org/codes/
relation: exactMatch
date: '2018-06-12'
specifiedBy:holds a Method object describing guidelines followed in generating the knowledge reported in the Statement
type: Method
name: ClinGen Hearing Loss Expert Panel Specifications to the ACMG/AMP Variant Interpretation Guidelines
reportedIn:a document that describes the Method
type: Document
urls:
- https://clinicalgenome.org/docs/clingen-hearing-loss-expert-panel-specifications-to-the-acmg-amp-variant-interpretation-guidelines/
hasEvidenceLines:holds EvidenceLine objects describing how difference types of evidence was interpreted to support the root Statement
- id: ex:EvidenceLine001an Evidence Line based on cohort allele frequency data from gnomAD (https://gnomad.broadinstitute.org/)
type: EvidenceLineuses the core EvidenceLine class as its type, but validated against the VariantPathogenicityEvidenceLine Profile
targetProposition: ex:Proposition001the possible fact against which evidence information is assessed in this EvidenceLine (typically, as here, this is the same proposition as asserted in the root Statement it supports)
hasEvidenceItems:the information interpreted as evidence in building this Evidence Line
- id: ex:StudyResult001here, the evidence consists of a single StudyResult, which collects several allele frequency data items about the 1-10120-T-G allele.
type: CohortAlleleFrequencyStudyResult
name: Overall Cohort Allele Frequency for 1-40819444_40819446-del
focusAllele: ex:Variant001the KCNQ4 variant that data included in this Result are about (the full representation of the variant is not included)
focusAlleleFrequency: 0
focusAlleleCount: 0three specific data items produced by the analysis are collected in this StudyResult (focus allele frequency, focus allele count, and locus allele count)
locusAlleleCount: 34086
sourceDataSet:the gnomAD dataset from which the data included in this Result were pulled.
id: gnomad4.1.0
type: DataSet
name: gnomAD v4.1.0
version: 4.1.0
cohort:a description of the cohort within the gnomad dataset interrogated in the analysis (here, the full gnomad population)
id: ALL
name: Overall
type: StudyGroup
specifiedBy:holds a Method object describing protocols and guidelines followed in generating the data reported in the Study Result
type: Method
name: gnomAD methods
reportedIn:a document that describes the Method (this is all we are given about this Method in the source data)
type: Document
name: gnomAD help documentation
urls:
- "https://gnomad.broadinstitute.org/help"
directionOfEvidenceProvided: supportsreports that the frequency evidence 'supports' the target proposition (as opposed to disputing it)
strengthOfEvidenceProvided:
primaryCoding:
code: moderatereports that this supporting evidence is of 'moderate' strength
system: ACMG Guidelines, 2015
evidenceOutcome:holds a single term summarizing evidence direction and strength assessments, using community-specific vocabulary ...
primaryCoding:
code: PM2_moderate... here, that the evidence line provides moderate evidence for Pathogenicity, based on the ACMG PM2 criteria
system: ACMG Guidelines, 2015
name: ACMG 2015 PM2 Moderate Criterion Met
specifiedBy:holds a Method object describing guidelines followed in generating the evidence assessment in this Evidence Line
type: Method
methodType: PM2
name: ClinGen Hearing Loss Expert Panel Specifications to the ACMG/AMP Variant Interpretation Guidelines
reportedIn:a document that describes the Method (this is all we are given about this Method in the source data)
type: Document
urls:
- https://clinicalgenome.org/docs/clingen-hearing-loss-expert-panel-specifications-to-the-acmg-amp-variant-interpretation-guidelines/
contributions:holds descriptions of contributions to this Evidence Line
- type: Contribution
contributor:
id: curator001
type: Agent
activityType:
name: evidence evaluation
date: '2018-03-11'
- id: ex:EvidenceLine002an Evidence Line based on functional impact data about the variant from MAVE (https://mavedb.org/)
type: EvidenceLineuses the core EvidenceLine class as its type, but validated against the VariantPathogenicityEvidenceLine Profile
targetProposition: ex:Proposition001
hasEvidenceItems:
- id: ex:Statement002here the evidence item is another Statement about the functional impact of the variant
type: Statement
proposition:
type: ExperimentalVariantFunctionalImpactProposition
subjectVariant: ex:Variant001the full representation of the variant subject of this Statement is not included
predicate: impactsFunctionOfthe predicate for this type of Statement is fixed at 'impactsFunctionOf'
objectSequenceFeature:holds a MappableConcept object that represents the Gene impacted by the variant, using names/codes from existing code systems
id: clinvar-gene:9132
conceptType: Gene
primaryCoding:
code: ncbigene:9132
system: https://identifiers.org/ncbigene
iris:
- https://identifiers.org/ncbigene:9132
name: KCNQ4
experimentalContextQualifier:this qualifier is able to take a custom, data provider-defined object to describe the experiment in which the reported impact was determined. A condensed example is shown here, but an real and complete example can be found in the Exp-Var-Func-Impact-Statement-01.yaml test fixtures file.
title: KCNQ4 VAMP Seq Expt 001
description: Multiplex assessment of KCNQ4 protein variant abundance by massively parallel sequencing
phenotypicAssay: flow cytometry
modelSystem: immortalized human cells
variantLibrarySystem: oligo-directed mutagenic PCR
profilingStrategy: barcode sequencing
sequencingReadType: single-segment (short read)
direction: supportsindicates that the Statement supports the assessed impact Proposition above (i.e. says that the subject Variant does impact the function of the object Gene)
classification:summarizes the Statement in terms of a final classification of the variant, using a term familiar in the community of use.
primaryCoding:
code: abnormalindicates the variant version of the gene has abnormal function (consistent with the 'impactsFunctionOf' proposition being 'supported')
system: ga4gh-gks-term:experimental-var-func-impact-classification
specifiedBy:a Method followed to produce the Statement, which is described by the publication indicated below
type: Method
methodType:
name: variant interpretation guideline
reportedIn:
type: Document
pmid: 29785012
hasEvidenceLines:
id: EvidenceLine003
type: EvidenceLine
directionOfEvidenceProvided: supportsindicates that EvidenceLine003 based on a Functional Impact Study Result 'supports' the Functional Impact Statement
specifiedBy:a Method followed in assessing the direction and strength of evidence provided by the Functional Impact StudyResult for the Functional Impact Statement
type: Method
name: MAVE bayesian threshold probability method 001
reportedIn:
type: Document
urls:
- "https://mavedb.org/score-sets/urn:mavedb:00000013-a-1"
hasEvidenceItems:a Study Result that captures the experimental data and scores on which the Functional Impact Statement was based.
- id: ex:StudyResult002the evidence in this case is data captured in a Functional Impact Study Result
type: ExperimentalVariantFunctionalImpactStudyResult
focusVariant: ex:Variant001the KCNQ4 variant that data are about (a full representation of the variant is not included)
functionalImpactScore: 1.29395467005388this is the only data item included right now in this StudyResult
specifiedBy:
type: Method
methodType:
name: Experimental protocol
reportedIn:
type: Document
pmid: 29785012
sourceDataSet:
type: DataSet
name: variant effect data set
license:
primaryCoding:
code: CC0
system: https://spdx.org/licenses/
iris:
- https://spdx.org/licenses/CC0-1.0.html
reportedIn:
type: Document
urls:
- "https://mavedb.org/score-sets/urn:mavedb:00000013-a-1"
directionOfEvidenceProvided: supportsindicates that EvidenceLine002 based on a Functional Impact Statement 'supports' the root Pathogenicity Statement
strengthOfEvidenceProvided:
primaryCoding:
code: strongindicates that this line of evidence provides 'strong' support for the variant's Pathogencity
system: ACMG Guidelines, 2015
evidenceOutcome:
primaryCoding:
code: PS3_strong
system: ACMG Guidelines, 2015
name: ACMG 2015 PS3 Supporting Criterion Met
specifiedBy:holds a Method object describing guidelines followed in assessing the evidence provided by the Functional Impact Statement for the root Pathogenicity Statement
type: Method
methodType: PS3
name: ClinGen Hearing Loss Expert Panel Specifications to the ACMG/AMP Variant Interpretation Guidelines
reportedIn:
type: Document
urls:
- https://clinicalgenome.org/docs/clingen-hearing-loss-expert-panel-specifications-to-the-acmg-amp-variant-interpretation-guidelines/
contributions:
- type: Contribution
contributor:
id: curator002the curator who assessed functional impact statement as evidence for pathogenicity
type: Agent
activityType:
name: evidence evaluation
date: '2018-04-03'
extensions:holds Extension objects which allow data providers to define key-value pairs for capturing additional info not supported by the VA model.
- name: clinvarMethodCategoryhere, Extensions are used to report clinvar-specific values that the data provider does not want to lose
value: literature only
- name: clinvarReviewStatus
value: no assertion criteria provided
- name: clinvarSubmittedClassification
value: Pathogenic
Detailed Diagram:
The diagram shows a subset of data from the full json example. It provides a more detailed data structure overview that highlights encapsulation of Propositions in Statements and Evidence Lines and the use of the same set of Core Model classes (Method, Document, Contribution, Agent) to capture provenance information about all primary knowledge artifacts.
It also highlights the kind of schema that specifies each objects in the data - illustrating how Core Model Classes, Base Profiles, and Community Profiles that rely on different authoring mechanisms are used together in a structured data representation.
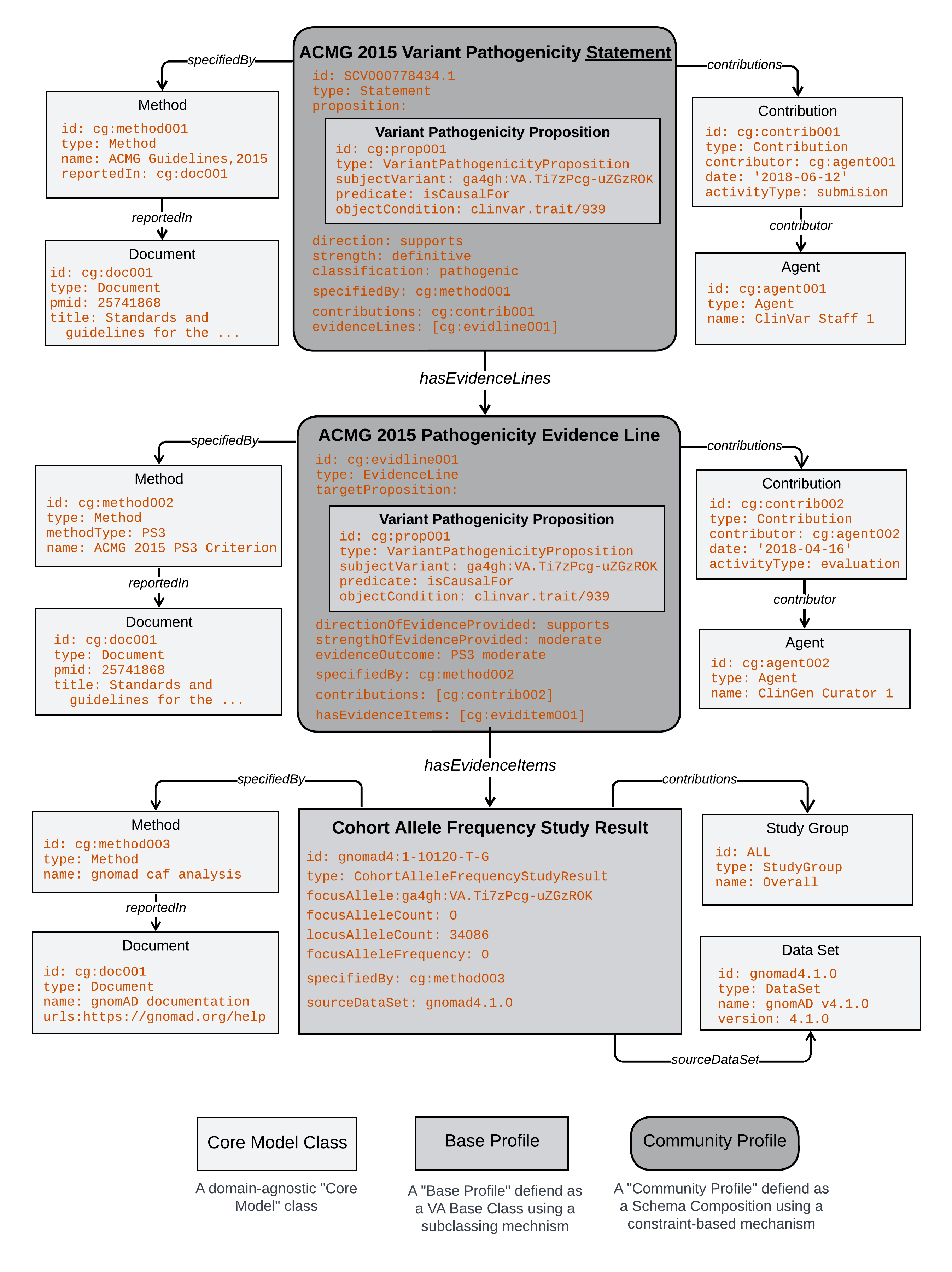
Detailed Data Example
Legend: Diagrammatic representation of a subset of data in the json example above. Styling conventions indicate the type of model that specifies each object in the example (Core Class, Base Profile, Community Profile). To fit the data into this form and make it human readable, syntactic shortcuts were taken to simplify values normally wrapped in complex data structures like MappableConcepts and Codings.
A key thing to note in the example is that, because Base Profiles are defined as formal subclasses, these objects have a specific type that reflects this (e.g. CohortAlleleFrequencyStudyResult). But because Community Profiles are defined using schema composition, the formal type of these objects is that of the Core Model class on which they are built (e.g. Statement, EvidenceLine).