Modeling Foundations
This document describes modeling standards, patterns, and principles employed by the VA Specification for representing genetic variation and knowledge about them.
After a brief overview of standards used to represent genetic variation, it focuses on data structures that can be built around three key classes which are the basis of VA Profiles: (1) Statements; (2) Study Results; and (3) Evidence Lines.
Variant Representation
To represent genetic variations that are subjects of VA Statements, the VA-Spec adopts two complementary GKS standards:
The GA4GH Variant Representation Specification (VRS), which provides JSON Schema for representing many classes of discrete genetic variation, and tools for generating globally-unique computed variant identifiers. VRS variants represent discrete instances of sequence variation in a specified context (reference, location, state) - e.g. the NM_005228.5(EGFR):c.2232_2250del(p.Lys745fs) variant here. This includes single continuous alleles, haplotypes, genotypes, and copy number changes.
The GA4GH Categorical Variation Representation Specification (Cat-VRS), which is built on top of VRS and provides a terminology and data model for describing ‘categorical’ variation concepts. Categorical variations are intensionally defined sets of variations, based on criteria that must be met for inclusion in a given category. Examples include BRAF V600 mutations and EGFR exon 19 deletions.
VRS and Cat-VRS models are directly imported for use in VA schema, and the VA-Spec reference implementation will incorporate VRS tools for identifier generation, normalization, and validation. See linked documentation above for more information about these specifications.
Statement Representation
In the VA-Spec data, each assertion of knowledge about a variant is captured in a self-contained Statement object. The VA Core-Model provides a rich and powerful structure where:
the Statement class roots a larger data structure supporting clear and precise tracking of the evidence and provenance information
the semantics of what is assessed or reported to be true in a Statement are explicitly structured in a Proposition object, in terms of subject, predicate, object, and qualifier attributes (SPOQ)
Statements can report a more nuanced assessment of the state of confidence or evidence surrounding this Proposition
These features of the CoreM odelStatement model are described below.
Statement Data Structure
In VA-Spec data, a Statement object roots the larger data structure below.

Statement Data Structure
Legend A class-level view of the Statement-based structures supported in VA-Spec data. Italicized text under class names illustrate the kind of information each class may report, in the case of a Variant Pathogenicity Statement supported by Population Allele Frequency evidence.
- In this structure:
A Statement object roots a central axis where it is linked to one or more Evidence Lines representing discrete arguments for or against it.
Each Evidence Line may then be linked to one or more Evidence Items - specific Information Entities that were used to build an evidence-based argument.
Surrounding this central axis are classes that describe the provenance of these artifacts, including Contributions made to them by Agents, Activities performed in doing so, Methods that specify their creation, and Documents that describe them.
This structure allows tracking of provenance information at the level of a Statement and each supporting Evidence Line and Item. A simple data example illustrating the structure for a Variant Pathogenicity Statement can be found here.
Statement Semantics
Every Statement object puts forth a Proposition - a possible fact it assesses or reports to be true - the semantics of which explicitly captured in a Proposition object using subject, predicate, object, and optional qualifier attributes (SPOQ).
An assessment of this Proposition’s validity can be captured using direction, strength, and score attributes (DS) - which indicate whether the Proposition is reported to be true or false, and the amount of confidence or evidence supporting this claim.
This “SPOQ-DS” model supports two “Modes of Use” for Statements, which differ in what they say about their Proposition, and can be distinguished by how the direction and strength or score attributes are populated.
In “Assertion Mode”, a Statement simply reports an SPOQ proposition to be true or false (e.g. that “BRCA2 c.8023A>G is pathogenic for Breast Cancer”). The
strength/scoreattributes are not populated, anddirectionis assumed to be true/supports if not otherwise indicated. This mode is used by project reporting conclusive assertions about a domain of discourse, but not providing confidence or evidence level assessments.In “Proposition Assessment Mode”, a Statement describes the overall state of evidence and/or confidence surrounding the SPOQ proposition - which is not necessarily being asserted as true. The
directionandstrength/scoreattributes are populated, which allows for more nuanced Statements reporting things like “there is weak evidence supporting the proposition that ‘BRCA2 c.8023A>G is causal for Breast Cancer’”, or “we have high confidence that the proposition ‘PAH:c.1285C>A is causal for Phenylketonuria is false”). This mode is used in projects to track the evolving state of support for propositions of interest, as curators actively collect evidence and work toward a conclusive assertion.
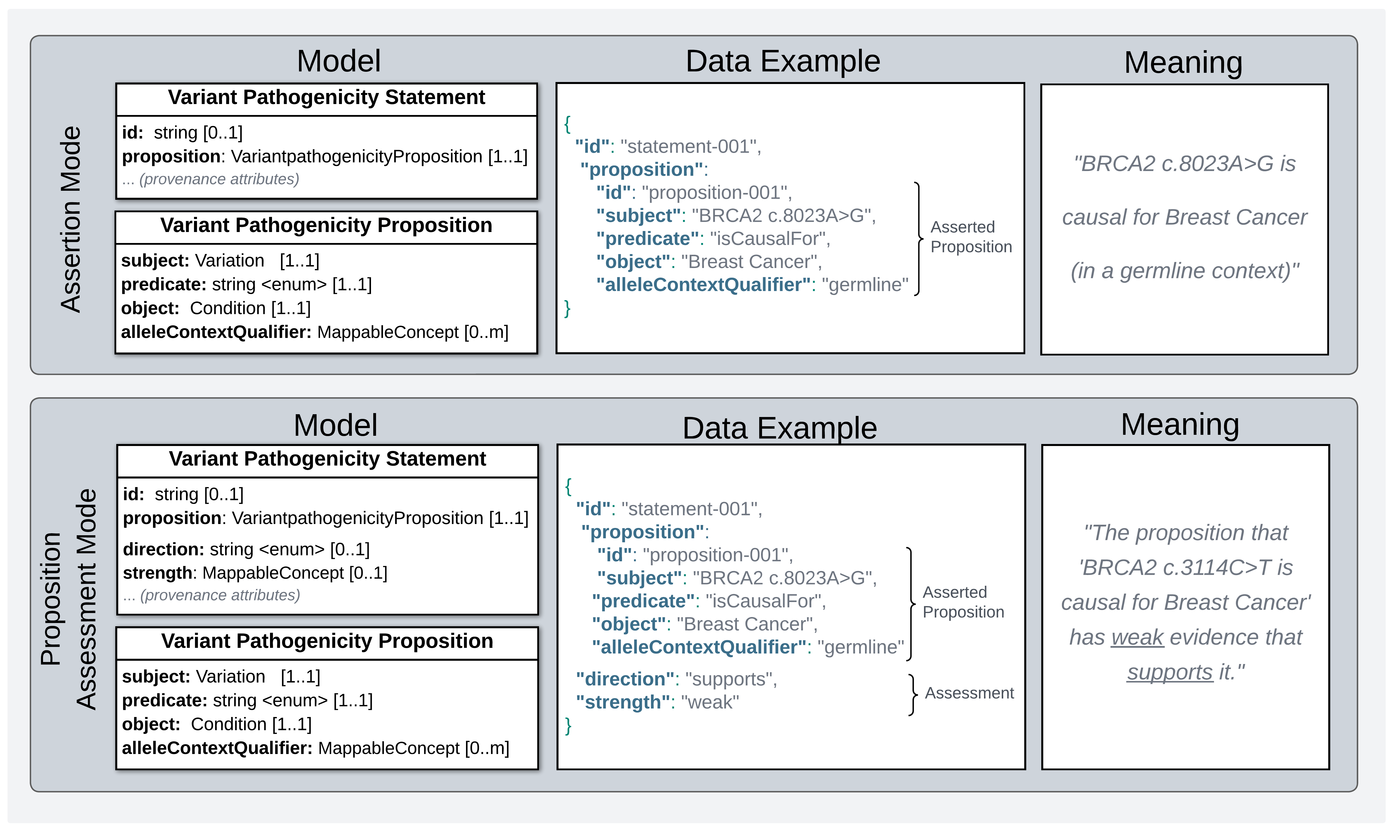
Statement Semantics in Two Modes of Use
Legend Assertion Mode vs Proposition Assessment Mode semantics for a Variant Pathogenicity Statement. Left Panels: Abridged versions of the data models for each mode of use, showing attributes reporting central claim each puts forth (most attributes describing evidence and provenance information are omitted). Center Panels: An example of a Variant Pathogenicity Statement instance. Right Panels: Plain language meaning of what structured data in the example reports to be true.
Implementations should choose the mode that best fits their data and use case when generating VA-compliant datasets - leveraging direction and strength/score attributes only if they wish to describe the state of evidence or confidence surrounding a possible fact.
Study Result Representation
Many users of the VA-Spec provide curated collections of data about a particular variant from a particular study or analysis, as opposed to higher order statements of knowledge. The GKS Core-IM defines the Study Result class to support this use case.
Like the Statement class, it roots a larger data structure supporting clear and precise tracking of evidence and provenance information, and provides explicit semantics linking a variant to specific data and study context. These features of the Core-IM Study Result model are described below.
Study Result Data Structure
In VA-Spec data, a Study Result object roots the data structure below:

Study Result Data Structure
Legend A class-level view of the Study Result-based structures supported in VA-Spec data. Italicized text under class names illustrate the kind of information each class may report in the case of a Cohort Allele Frequency Study Result reporting data from the gnomAD dataset about a particular variant.
In this structure, the data items collected in the Study Result can be linked to the larger Data Set or sets from which they came, and a description of the Study Group from which the data was collected. And as with Statements, clear and precise provenance information about the Study Result and DataSet can be captured in supporting Method, Document, Contribution, Agent, and Activity objects.
Study Result Semantics
Study Results simply report a collection of one or more data items from a particular study, that are presented together because they are about a common ‘focus’ (e.g. a particular variant), and typically produced by a common process or methodology.
Key attributes expressing core StudyResult semantics include:
A
focusattribute that captures the entity of interest that the data items are aboutA set of one or more data-type specific attributes created during the profile process, that capture the specific data items that are about this focus.
A
studyGroupattribute that reports information about the group of subjects that were interrogated to produce this data.A
sourceDataSetattribute that reports the larger data set from which selected data items came.
Additional attributes allow provenance information about the evidence assessment process and the underlying evidence to be captured (who created the data, when, using what methods, etc).
The diagram below highlights these key attributes, and provides an example of how this class structures allele count and frequency data about a particular variant in an east asian population from a gnomad study dataset.
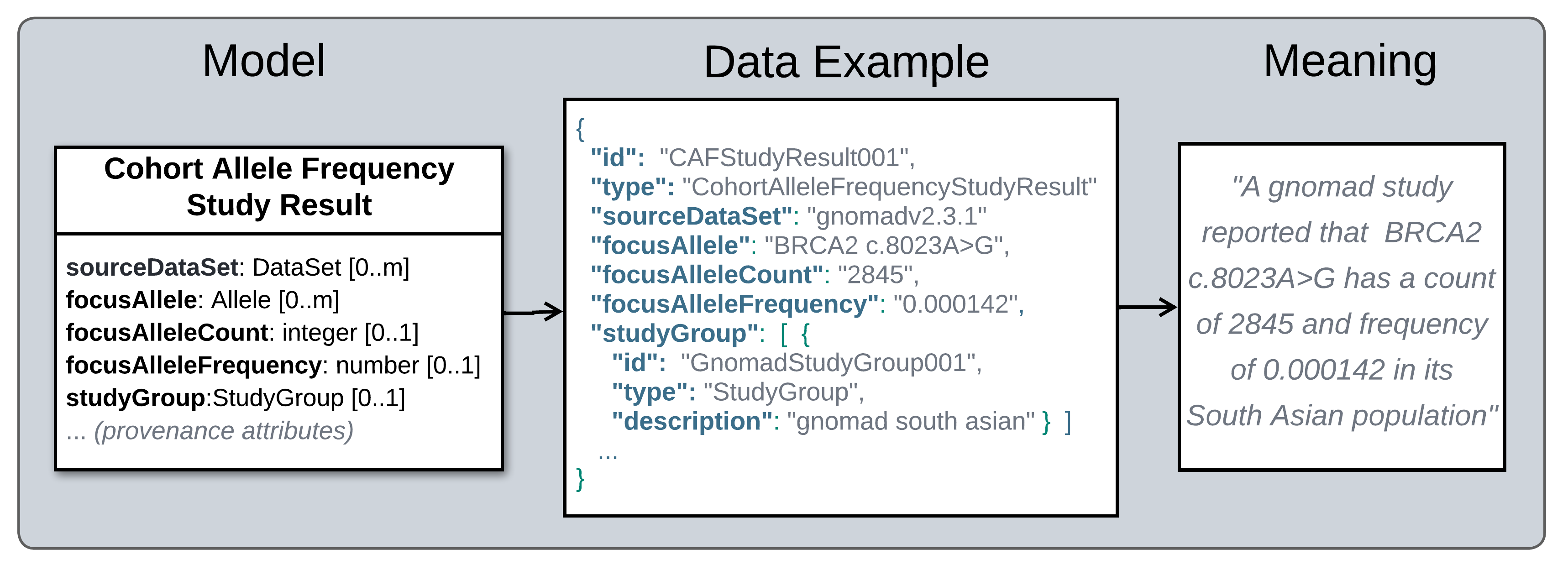

Semantic Meaning of Study Results.
Legend Semantics of a Cohort Allele Frequency Study Result. Left: An abridged version of a Cohort Allele Frequency (CAF) Study Result profile model, showing only attributes reporting the central claim it puts forth (attributes providing evidence and provenance information are omitted). Center: An example of a CAF Study Result instance. Right: Plain language meaning of structured data in the example.
Evidence Line Representation
Some users of the VA-Spec want to represent curated lines of evidence that describe how a collection of information (e.g. results from a published study, or subset of data items from a larger dataset) are interpreted as evidence supporting or disputing a possible fact that they ultimately want to be able to assert about a variant.
For example, that some set of allele frequency data items from gnomAD represents a moderate argument supporting the pathogenicity of a particular variant for a particular disease. Many organizations ‘pre-curate’ such arguments so that they can be tracked and efficiently combined to support assertions once sufficient evidence exists - e.g. that a variant is definitively pathogenic for the disease.
The GKS Core-IM defines the Evidence Line class to support this use case. Like the Statement class, it roots a larger data structure supporting clear and precise tracking of provenance information, and provides explicit semantics about nature of the argument being reported. These features of the Core-IM Evidence Line model are described below.
Evidence Line Data Structure
In VA-Spec data, an Evidence Line object can root the data structure below:

Evidence Line Data Structure
Legend A class-level view of the Evidence Line-based structures supported in VA-Spec data. Italicized text under class names illustrate the kind of information each class may report - here for an Evidence Line representing a moderate argument supporting the pathogenicity of a particular variant, based on allele frequency data from gnomAD.
In this structure, the Evidence Items contributing to the Evidence Line can be grouped and tied to an assessment of the direction and strength of support provided for or against a particular ‘Target Proposition’ (the possible fact towards which the evidence is assessed). And as with Statements, clear and precise provenance information about the Evidence Line and Evidence Items can be captured in supporting Method, Document, Contribution, Agent, and Activity objects.
Evidence Line Semantics
Evidence Lines represent a type of information that sits between foundational evidence and a final assertion of purported fact, in the process of generating scientific knowledge. Here, they express the idea that a particular collection of evidence items was interpreted to provide a particular strength and direction of support for or against the possible fact expressed in this asserted Statement. Is is through the collective assessment of one or more lines of evidence that hypothesis and conjecture become cemented as scientific fact.
The attributes defined in the Evidence Line class are carefully crafted to express these core elements of an Evidence Line:
The
evidenceItemsattribute captures the information that was assessed as evidence.The
targetPropositionattribute reports the ‘possible fact’ that the evidence is assessed against.The
directionOfEvidenceProvidedandstrengthOfEvidenceProvidedattributes report the outcome of this assessment - whether the evidence line supports or disputes the target proposition, and how strongly.
Additional attributes allow provenance information about the evidence assessment process and the underlying evidence to be captured (who did it, when, using what guidelines, etc).
The diagram below highlights key Evidence Line attributes, and provides an example of how this class structures data to report that the allele count and frequency data in a particular cohort allele frequency study result provides moderate evidence supporting the Pathogenicity of BRCA2 c.8023A>G.
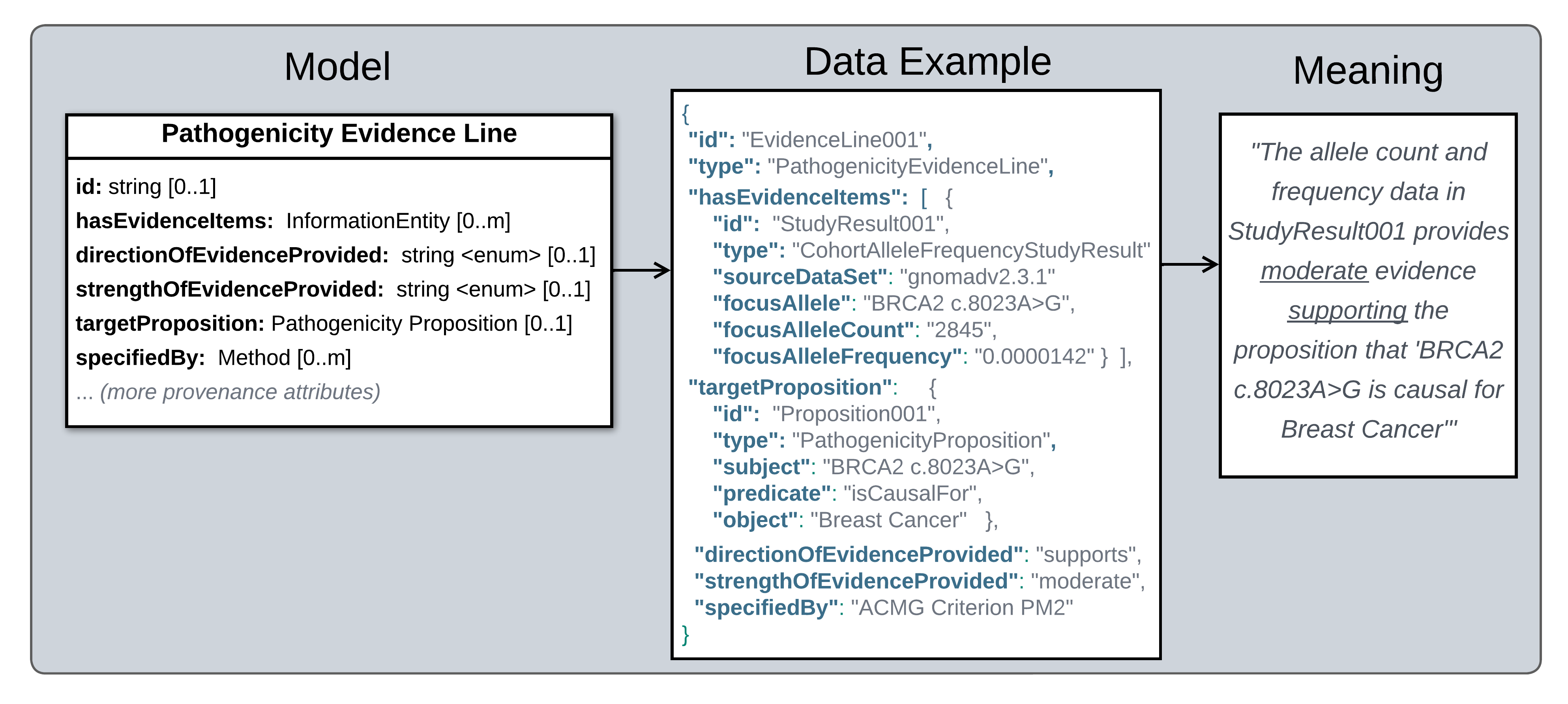
Semantic Meaning of Evidence Lines.
Legend Semantics of a Pathogenicity Evidence Line. Left: An abridged version of a Pathogenicity Evidence Line profile model, showing only attributes reporting the central claim it puts forth (attributes describing provenance information are omitted). Center: An example of a Pathogenicity Evidence Line instance. Right: Plain language meaning of the structured data in the example.